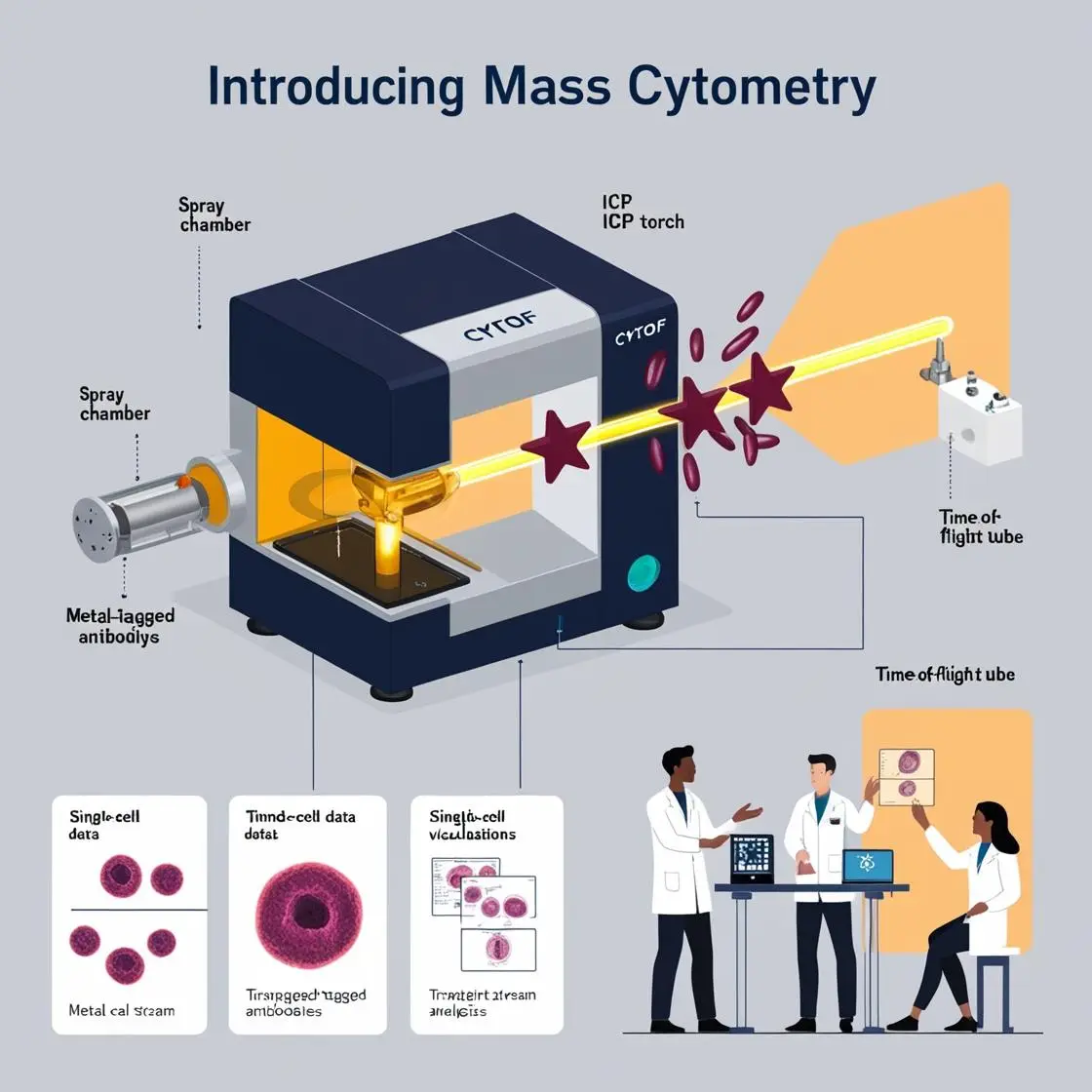
The advent of spatial transcriptomics and proteomics has ushered in a new era of biological understanding, where the location of cells within tissues is as important as their molecular profiles. This chapter explores the exciting intersection of imaging mass cytometry with spatial transcriptomics, offering unprecedented insights into tissue architecture and cellular interactions.
Single-Cell Spatial Analysis in Situ
The ability to perform single-cell analysis within the native tissue environment has opened new avenues for understanding cellular behavior and interactions.
A landmark study by Keren et al. (2018) in Cell, cited over 830 times, “A Structured Tumor-Immune Microenvironment in Triple Negative Breast Cancer Revealed by Multiplexed Ion Beam Imaging,” used highly multiplexed ion beam imaging to analyze breast cancer samples at single-cell resolution. This work revealed complex spatial relationships between tumor and immune cells, providing insights into tumor progression and potential therapeutic targets. It is actually another version of the CyTOF, but where spatial resolution has been permitted.
Combining Imaging Mass Cytometry with Spatial Transcriptomics
The integration of imaging mass cytometry (IMC) with spatial transcriptomics represents a powerful approach to understanding cellular heterogeneity within the context of tissue architecture. This combination allows for the simultaneous visualization of protein expression and gene transcription at single-cell resolution within intact tissues.
A groundbreaking study by Goltsev et al. (2018), cited over 1100 times and published in Cell, “Deep profiling of mouse splenic architecture with CODEX multiplexed imaging,” demonstrated the power of this approach. Using CODEX (CO-Detection by indEXing) technology, they created a high-dimensional map of mouse spleen at single-cell resolution, revealing complex cellular neighborhoods and previously unknown cell types.
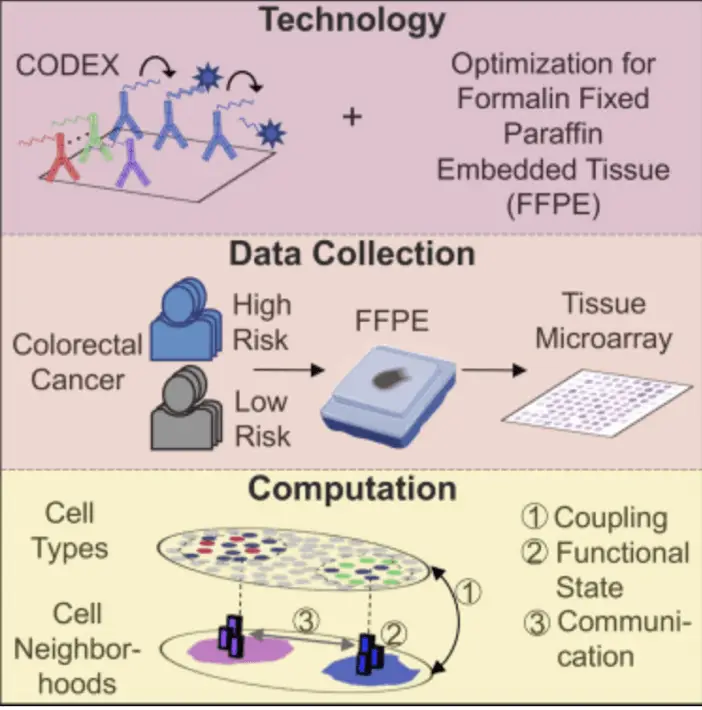
Building on this foundation, Schürch et al. (2020) in Cell, cited over 600 times, “Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front,” combined IMC with spatial transcriptomics to dissect the tumor microenvironment in colorectal cancer. This study revealed how specific cellular neighborhoods influence anti-tumor immune responses, providing new insights for cancer immunotherapy.
This image presents a novel approach to analyzing colorectal cancer risk using advanced tissue analysis techniques. It combines CODEX imaging technology, optimized for FFPE samples, with sophisticated computational analysis. The method collects data from high and low-risk colorectal cancer patients, using tissue microarrays for high-throughput analysis. The computational approach focuses on cell coupling, functional states, and intercellular communication, providing a comprehensive view of the tumor microenvironment. Unlike imaging CyTOF, this CODEX-based method offers higher spatial resolution, less-destructive imaging, and is more tailored for clinical applications due to its FFPE optimization. The approach’s novelty lies in its integrative analysis, preservation of spatial information, multi-parameter examination, and direct clinical relevance. By offering a more comprehensive, spatially-resolved analysis of tumor tissues, this method could significantly advance cancer research, potentially improving risk stratification and identifying new therapeutic targets based on cell-cell interactions in the tumor microenvironment.
3D Reconstruction of Tissue Architecture
The ability to reconstruct tissue architecture in three dimensions adds another layer of complexity to our understanding of cellular organization. Advanced imaging techniques combined with computational approaches are making this possible.
In the realm of proteomics, Ijsselsteijn et al. (2021), working also at the LUMC, published in Frontiers, “A 40-marker panel for high dimensional characterization of cancer immune microenvironments by imaging mass cytometry”
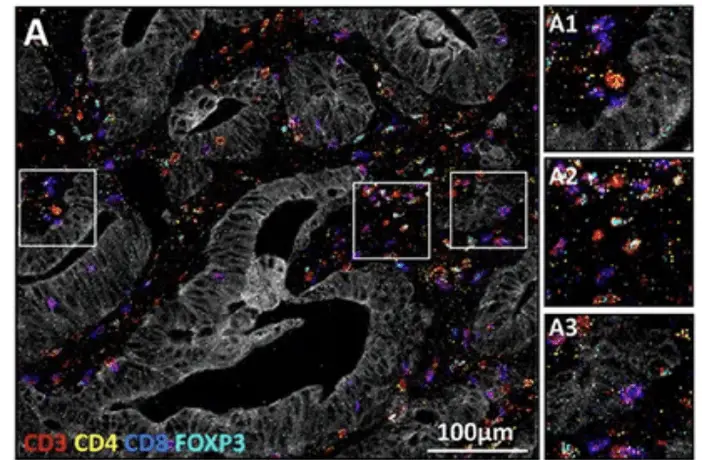
Later, a pioneering study by Wang et al. (2022) in Science, cited over 1100 times, “Three-dimensional intact-tissue sequencing of single-cell transcriptional states,” introduced a method for 3D spatial transcriptomics in intact tissue samples. This technique, called STARmap3D, allows for the visualization of gene expression patterns in three dimensions, providing insights into complex tissue organizations.
The Power of Spatial Context
The integration of spatial information with molecular profiles is revolutionizing our understanding of biology. From unraveling the complexities of the tumor microenvironment to mapping the intricate architecture of healthy tissues, these technologies are providing unprecedented insights.
For instance, in neuroscience, spatial transcriptomics is helping to create detailed maps of brain regions, revealing how gene expression patterns relate to neuronal function and connectivity. In developmental biology, these techniques are shedding light on the spatial cues that guide organ formation and cellular differentiation.
The ability to view tissues through this high-dimensional, spatially resolved lens is not just an academic exercise. It has profound implications for medicine. In cancer research, understanding the spatial relationships between tumor cells and immune cells is guiding the development of more effective immunotherapies. In autoimmune diseases, mapping the distribution of immune cell subsets within affected tissues is providing new insights into disease mechanisms and potential treatment targets.
As we continue to refine these technologies and develop new computational tools for data analysis, we are uncovering the hidden languages of cellular communication written in the spatial organization of tissues. Each study adds a new layer to our understanding, revealing the intricate tapestry of life at a resolution previously unimaginable.
The field of spatial transcriptomics and proteomics stands at the forefront of a new era in biology, where the where is as important as the what. As we decode the spatial syntax of tissues, we move closer to a holistic understanding of biological systems, promising new insights into health and disease. This journey of discovery, mapping the cellular landscapes of life, is transforming our view of biology from flat snapshots to rich, multidimensional portraits, each telling a unique story of cellular life and interaction.
Spatial cytometry is like looking at a cellular city from a helicopter - fascinating, but oh, is it complex! It's one thing to understand cells in suspension, but add in their spatial relationships and suddenly you're playing 3D chess while everyone else is still on checkers.
Guillaume Beyrend