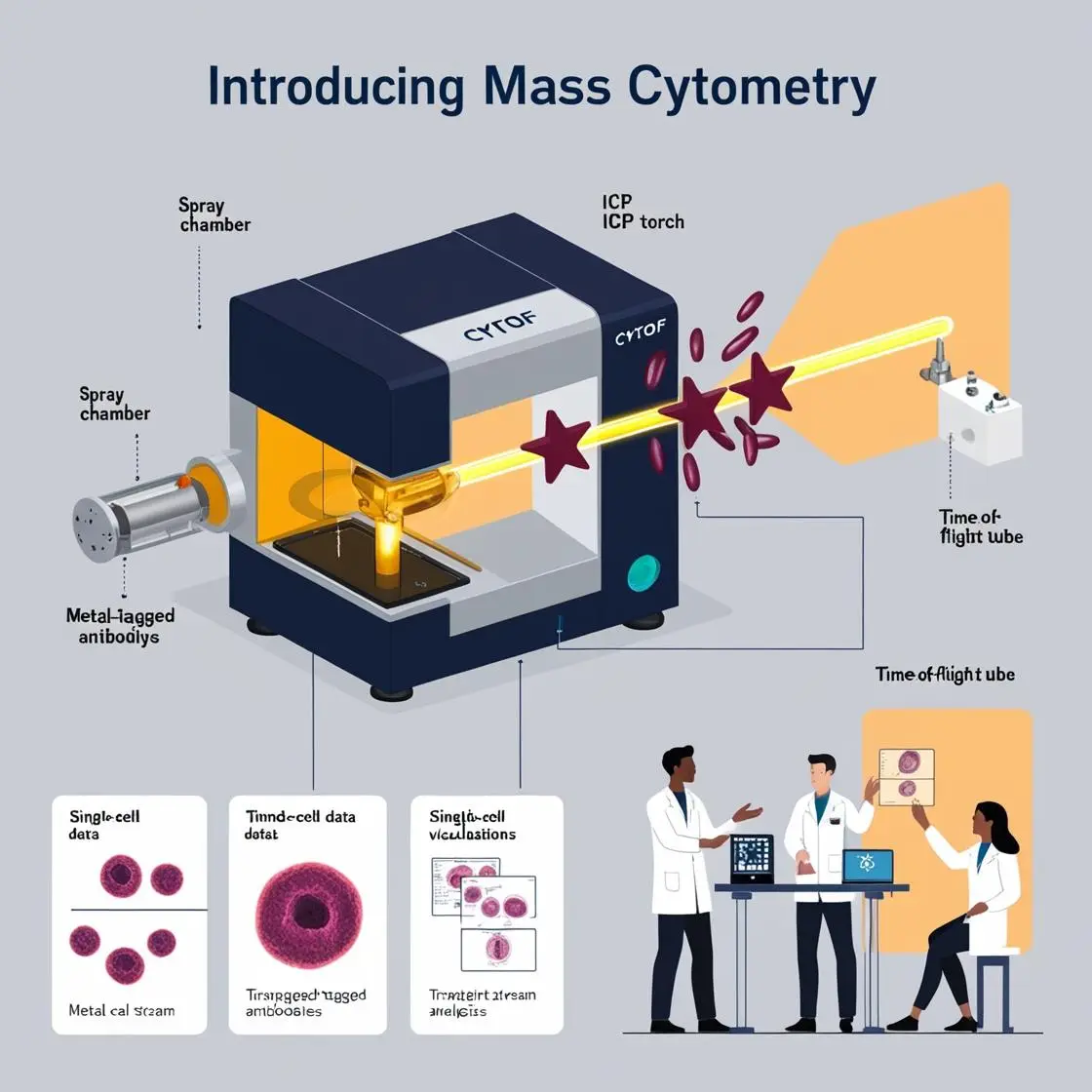
The art and science of designing mass cytometry panels are critical skills in the modern immunologist’s toolkit. As we delve into the intricacies of cellular systems, the ability to simultaneously measure dozens of parameters becomes both a powerful tool and a complex challenge.
Selecting Appropriate Markers
The first step in panel design is choosing the markers for your research question. This process requires a deep understanding of the biological system under study and the available reagents. As highlighted in the work of Takahashi et al. (2017), published in Cytometry Part A, the selection of markers should be guided by biological relevance and technical considerations.
Key considerations include:
- Biological relevance: Choose markers that are known to be important in your system of interest.
- Expression levels: Include a mix of high and low-expression markers to capture the full range of cellular phenotypes.
- Cellular localization: Include markers from different cellular compartments (surface, intracellular, nuclear) to view the cellular state comprehensively.
Avoiding Signal Overlap and Spillover
One key advantage of mass cytometry over traditional flow cytometry is the minimal signal overlap between channels. However, some spillover can still occur, particularly with high-abundance proteins or channels with similar masses.
Takahashi et al. (2017) proposed a novel approach to panel design that considers potential signal interference. Their “designed distribution of signal interference” method involves deliberately distributing potential interference across the panel to minimize its impact on any measurement.
Optimizing Panel Design for Specific Research Questions
The ultimate goal of panel design is to maximize the information gained about your biological system of interest. This often involves iterative optimization and tailoring to specific research questions.
In our recent work (Beyrend et al., 2023), we employed a sophisticated 42-marker panel to dissect the complex interplay between T-cell costimulation and checkpoint blockade in cancer immunotherapy. Our study, “OX40 agonism enhances PD-L1 checkpoint blockade by shifting the cytotoxic T cell differentiation spectrum,” showcased how a carefully designed high-dimensional panel can reveal subtle but crucial changes in cellular phenotypes.
Pushing the Boundaries: High-Dimensional Panels and Data Integration
While our 42-marker panel represents a highly sophisticated approach, the field continues to push towards even higher dimensionality and more comprehensive analysis. A landmark study by Chevrier et al., published in Cell in 2018 and cited over 900 times, exemplifies this trend. In their work “An Immune Atlas of Clear Cell Renal Cell Carcinoma,” they utilized two separate high-dimensional panels: one for myeloid cells and another for T cells. Although these panels were not merged, they each provided deep insights into specific cellular compartments of the tumor microenvironment.
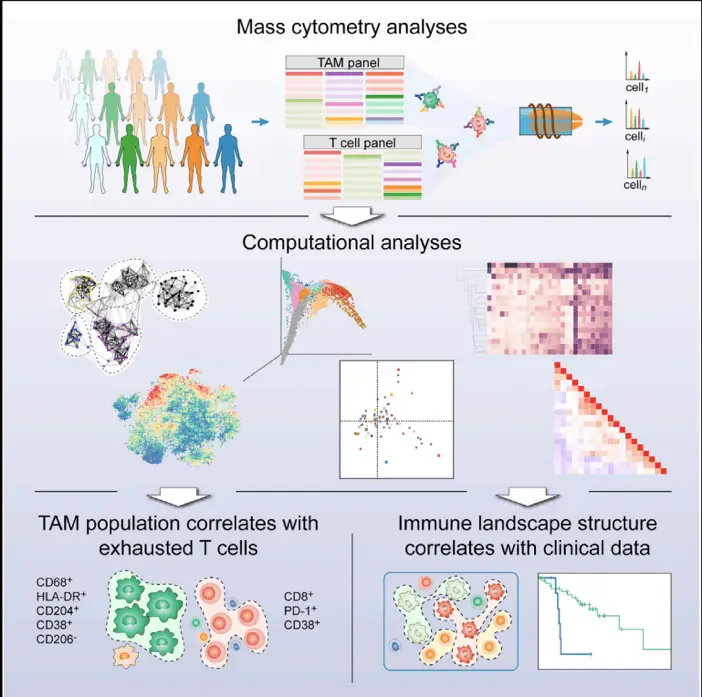
In the quest to push the boundaries of high-dimensional cytometry, researchers have developed innovative methods to expand the number of parameters that can be analyzed simultaneously. One such breakthrough came from our own lab with the development of “CYTOFmerge : integrating mass cytometry data across multiple panels”, Bioinformatics 2019. This computational tool allows for the seamless integration of data from two distinct CyTOF panels, effectively doubling the number of markers that can be analyzed in a single experiment. By identifying shared markers between panels and using them as anchors, CYTOFmerge aligns and merges datasets, creating a unified high-dimensional view of cellular phenotypes. This elegant solution not only addresses the upper limit of parameters in a single CyTOF run but also opens up new avenues for discovering subtle cellular states that might otherwise remain hidden. The ability to merge panels represents a significant leap forward in our capacity to unravel the complexities of cellular systems, paving the way for more comprehensive and nuanced analyses in fields ranging from immunology to cancer research.
CITE-seq: Bridging Proteomics and Transcriptomics
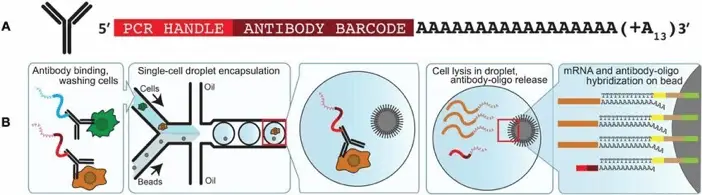
Another groundbreaking technique pushing the boundaries of single-cell analysis is CITE-seq (Cellular Indexing of Transcriptomes and Epitopes by Sequencing), as described in “Simultaneous epitope and transcriptome measurement in single cells”, Brief Communication, 2017. This method allows for simultaneous measurement of surface protein levels and mRNA in single cells.
In CITE-seq:
- Antibodies tagged with oligonucleotides (instead of metal isotopes) are used to label cell surface proteins.
- These antibody-derived tags are captured along with mRNA during single-cell RNA sequencing.
- This allows for combined protein and gene expression profiling at the single-cell level.
CITE-seq offers several advantages:
– It provides both proteomic and transcriptomic data from the same cell.
– It can potentially measure more parameters than mass cytometry alone.
– It allows the discovery of new cell types and states based on combined protein and gene expression profiles.
The field of high-dimensional single-cell analysis is rapidly evolving, with mass cytometry at its forefront. From carefully designed individual panels to computational integration of multiple panels and the emergence of hybrid techniques like CITE-seq, researchers have an unprecedented ability to probe the complexities of cellular systems.
These advancements in panel design, data integration, and multi-omic profiling are not just technical achievements; they represent powerful new tools in our quest to understand and combat disease at the cellular level. As we continue to push the boundaries of what’s possible with these technologies, we stand on the brink of discoveries that were unimaginable just a few years ago. However, CITE-seq’s protein detection sensitivity is generally lower than that of flow cytometry or mass cytometry. This is partly due to the indirect nature of protein quantification through DNA-barcoded antibodies.
CITE-seq can be expensive. As of 2023, the cost per cell for CITE-seq analysis can range from $0.50 to $2, depending on the depth of sequencing and the number of protein markers.
I have quite suffered during my own PhD from the joys of panel design. Let me tell you, crafting the perfect mass cytometry panel is like trying to organize a dinner party for a hundred people with wildly different dietary requirements – in a tiny apartment. For four long months, I wrestled with this beast, armed with nothing but an A3-sized sheet of paper and an overworked pencil. This sheet became my constant companion, my confidant, my therapist. It started off crisp and white, full of hope and possibility. By the end, it was yellowed and wrinkled, a battle-scarred veteran of countless brainstorming sessions and midnight epiphanies. I couldn't bring myself to throw it away. It was like my own personal Declaration of Independence, if the Founding Fathers had been really into antibodies and metal isotopes. Now, of course, there's an algorithm that can do all this in a fraction of the time. CATALYST, they call it, probably because "Stealer of Scientists' Best Anecdotes" was too long. But you know what? I wouldn't trade my yellow notepad for all the algorithms in Silicon Valley. After all, what's a good scientific breakthrough without a story of perseverance, late nights, and a paper product that's seen better days? Sometimes, the best catalyst for discovery is good old-fashioned elbow grease – and a very patient writing pad.
Guillaume Beyrend