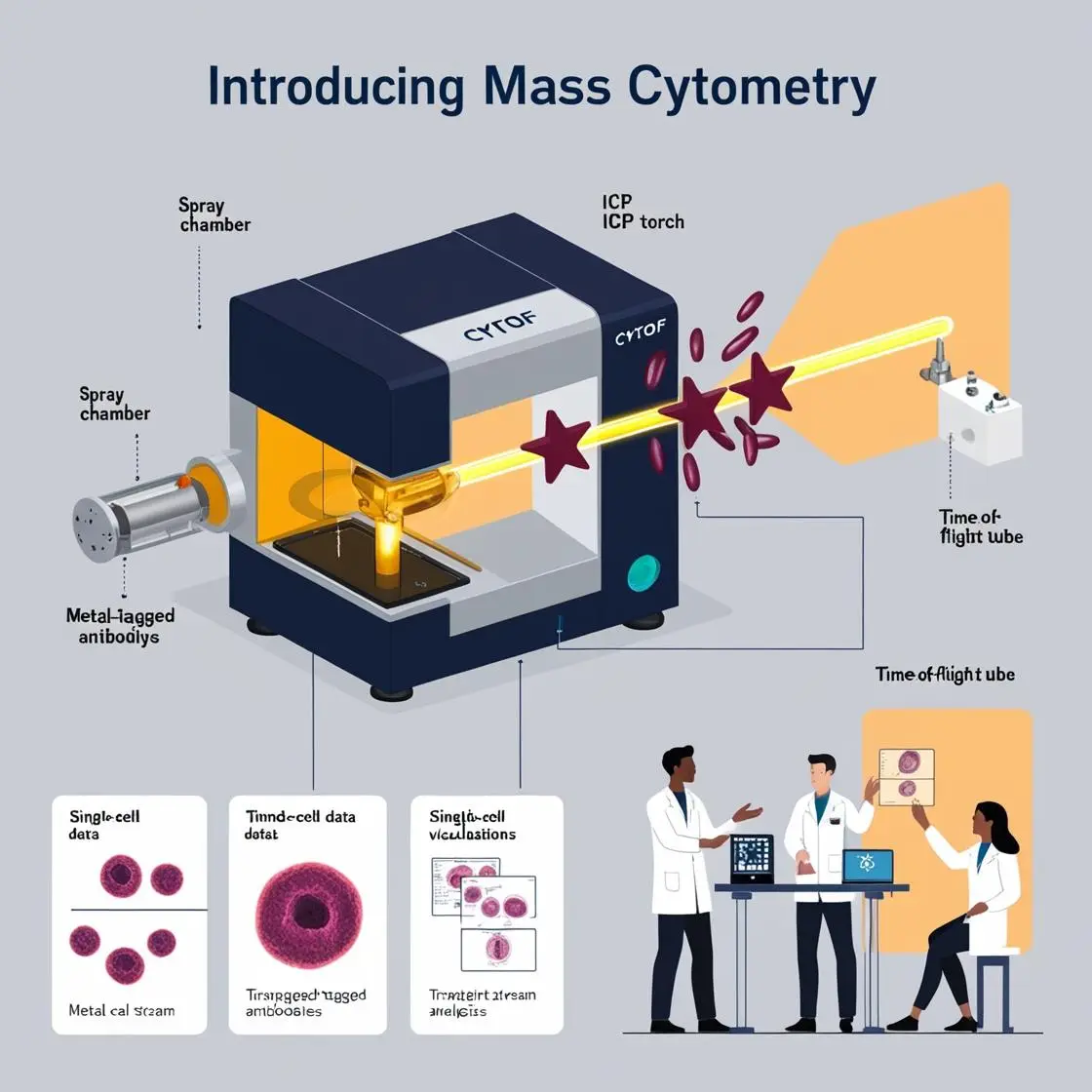
The integration of mass cytometry with other omics technologies represents a frontier in biomedical research, promising to provide a more comprehensive understanding of cellular biology. This chapter explores the exciting developments in multi-omics approaches, with a focus on how mass cytometry data can be integrated with other high-dimensional datasets.
Multi-omics Approaches Combining Mass Cytometry
The power of integrating mass cytometry with other omics technologies was beautifully demonstrated in a study by Spitzer et al. (2015) published in Science, “An interactive reference framework for modeling a dynamic immune system” (Spitzer et al., 2015, Science, 349(6244), 1259425). This groundbreaking work combined CyTOF with single-cell RNA sequencing to create a comprehensive map of the immune system, revealing new cell states and developmental trajectories.
Building on this foundation, a more recent study by Frohlich et al. (2022) in Nature Biotechnology, “Scalable prediction of acute myeloid leukemia using high-dimensional machine learning and blood transcriptomics,” integrated CyTOF data with bulk RNA sequencing and clinical information to develop a powerful predictive model for acute myeloid leukemia outcomes (Frohlich et al., 2022, Nature Biotechnology, 40(11), 1650-1658).
As often, papers using CyTOF are most of the time also using single-cell RNA sequencing, since both are deeply analysing immune cells.
Single-cell Transcriptomics and Proteomics Integration
The integration of single-cell transcriptomics with proteomic data from mass cytometry has opened new avenues for understanding cellular heterogeneity. A landmark paper in this area is the work by Peterson et al. (2017) in Nature Methods cited over 800 times, “Multiplexed quantification of proteins and transcripts in single cells”. This study introduced REAP-seq, a method that allows simultaneous measurement of cellular proteins and mRNAs at the single-cell level, quite similar to CITE-seq. While antibodies used in CITE-seq are conjugated to streptavidin that is noncovalently bound to biotinylated DNA barcodes, REAP-seq relies on covalent bonds between the antibody and aminated DNA barcode.
More recently, Mimitou et al. (2021) published a study in Nature Biotechnology titled “Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells” (Mimitou et al., 2021, Nature Biotechnology, 39(10), 1246-1258). This work introduced DOGMA-seq, which integrates single-cell ATAC-seq, RNA-seq, and protein quantification, providing a multi-omic view of cellular states.
Computational Methods for Data Integration
The integration of diverse omics datasets presents significant computational challenges. The review by Stuart and Satija (2019) in Nature Reviews Genetics provides an excellent overview of the computational methods developed for integrative single-cell analysis (Stuart & Satija, 2019, Nature Reviews Genetics, 20(5), 257-272).
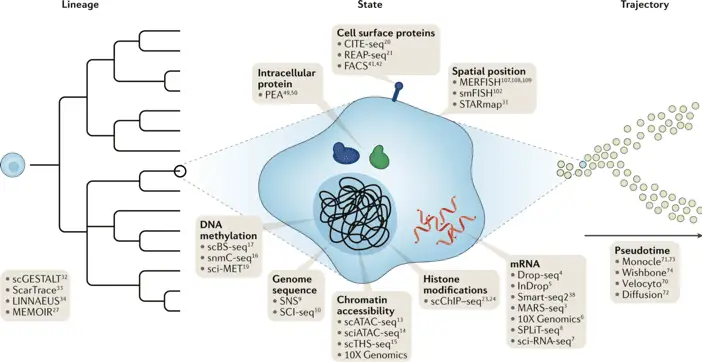
Integrating Metabolomics and Epigenetics
In the realm of epigenetics, Kelsey et al. (2018) in Nature Communications, cited over 600 times, “scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells,” introduced a method to simultaneously profile DNA methylation, chromatin accessibility, and gene expression in single cells (Kelsey et al., 2022, Nature Methods, 19(7), 823-832).
A particularly elegant example of data integration comes from the work of Argelaguet et al. (2020) published in Molecular Systems Biology and cited over 470 times, “MOFA+: a statistical framework for comprehensive integration of multi-modal single-cell data”. MOFA+ enables the integration of multiple molecular layers (e.g., RNA sequencing, DNA methylation, chromatin accessibility) at single-cell resolution.
Hartmann et al. (2021) in Cell, “Single-cell metabolic profiling of human cytotoxic T cells,” developed a method to combine mass cytometry with single-cell metabolomics. This innovative approach allowed for the simultaneous measurement of metabolic states and protein expression in individual immune cells.
The Power of Integration: Recent Breakthroughs
Several recent studies exemplify the potential of multi-omics integration, as stated in their comprehensive reviEw “Multi-Omics Profiling for Health” by Baby et al. (2023), Molecule and Cell Proteomics and by another systemic review “Applications of multi-omics analysis in human diseases”
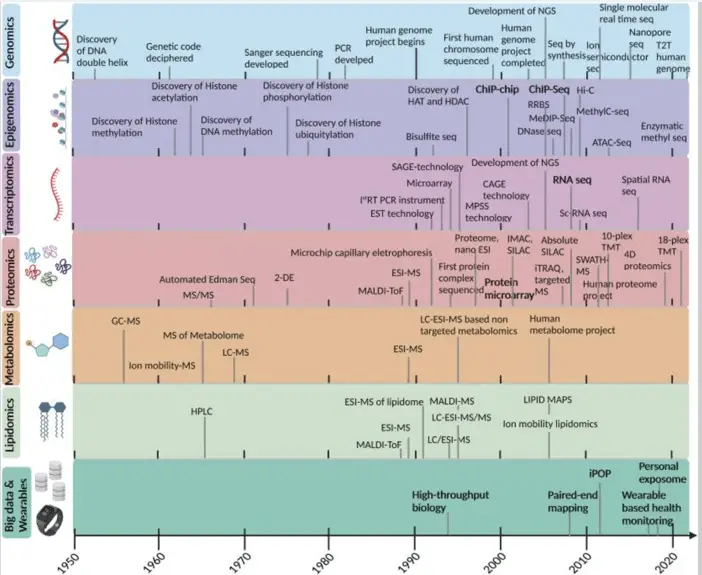
- Pekayvaz et al. (May 2024) in Nature Medicine, “Multiomic analyses uncover immunological signatures in acute and chronic coronary syndromes,” used MOFA technique to combine single-cell transcriptomics, epigenomics, and proteomics to uncover multicellular immune programs in cardiovascular disease, opening new directions for mechanistic, biomarker and therapeutic studies.
- Sameh et al. (2023) in Scientific Reports, “Integrated multiomics analysis to infer COVID-19 biological insights,” integrated CyTOF data with single-cell transcriptomics and serum antibody profiling to provide a comprehensive view of the humoral immune response in COVID-19.
- Weninger et al. (2023) in Scientific data “Single-cell integrative analysis reveals consensus cancer cell states and clinical relevance in breast cancer” reveal the association of immune-related state with better survival. Their study provides new insights for the cancer cell state composition and potential therapeutic strategies of breast cancer.
The Future of Multi-omics Integration
As we stand at the intersection of various omics technologies, we are witnessing the dawn of a new era in biological understanding. The integration of mass cytometry with transcriptomics, epigenomics, metabolomics, and other high-dimensional datasets is not just a technical achievement – it’s a conceptual leap that allows us to view cellular biology in its full complexity.
From slope-side epiphanies to cutting-edge studies, the field of multi-omics integration is rapidly evolving. Each integrated dataset is like a new lens through which we can view the cellular world, revealing patterns and relationships that were previously hidden from view.
As we continue to develop new methods for data integration and interpretation, we move closer to a holistic understanding of cellular biology. This integrated view promises to revolutionize our approach to complex biological questions, from understanding disease mechanisms to developing personalized therapies.
The journey from single-omics to multi-omics analysis mirrors our evolving understanding of biology itself – from a reductionist view of individual components to a systems-level appreciation of the intricate interplay between various cellular processes. As we ski down this exciting slope of scientific discovery, the view from the top is breathtaking, and the ride down promises to be exhilarating, offering unprecedented insights into the complex tapestry of life at the cellular level.
In our recent paper, "OX40 agonism enhances PD-L1 checkpoint blockade by shifting the cytotoxic T cell differentiation spectrum" (Beyrend et al., 2023), we took a deep dive into the world of multi-omics integration. We combined single-cell RNA sequencing, CyTOF, and functional data to paint a comprehensive picture of T cell responses to immunotherapy. The idea of merging these diverse omics technologies didn't strike me in a lab or during a late-night data analysis session. No, it hit me between ski runs at a Keystone conference in Breckenridge, Colorado, by seeing all these scientific talks. This experience taught me that combining technologies isn't just additive; it's multiplicative in terms of insights. However, it also requires a whole new level of skill to interpret the resulting data tsunami. It's like trying to conduct an orchestra where each instrument is from a different planet - challenging, but when it comes together, it's nothing short of magnificent.
Guillaume Beyrend