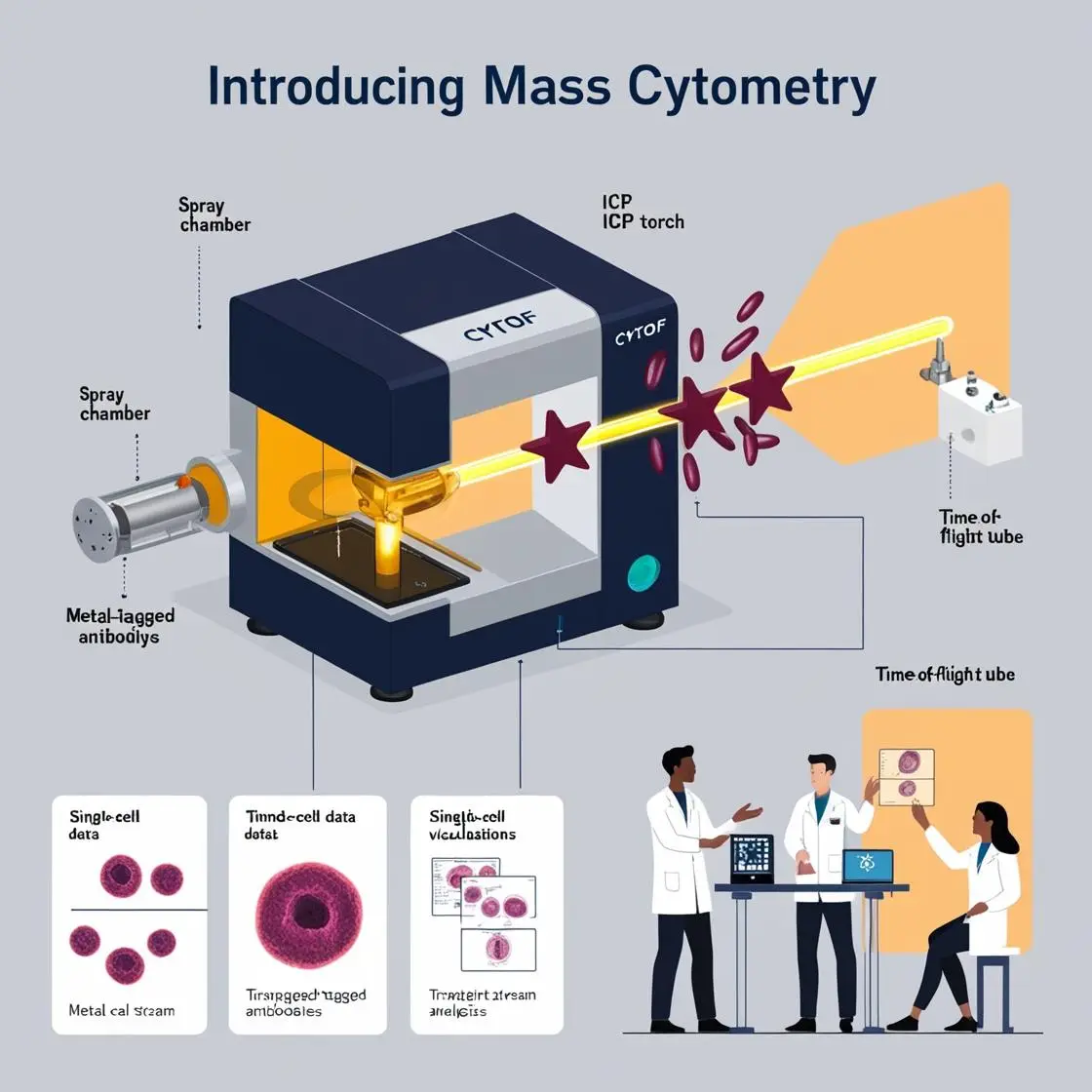
The integration of epigenetic analysis with mass cytometry has opened new frontiers in understanding cellular heterogeneity and gene regulation at the single-cell level. This chapter explores the exciting developments in combining these powerful technologies, with a focus on histone modification profiling, DNA methylation analysis, and applications in tumor immunology.
Combining Mass Cytometry with Epigenetic Analysis
The landmark study by Cheung et al. (2018) published in Cell demonstrated the power of combining mass cytometry with epigenetic profiling : “Single-Cell Chromatin Modification Profiling Reveals Increased Epigenetic Variations with Aging”. They developed a technique called EpiTOF (Epigenetic landscape profiling using cytometry by time-of-flight) to analyze 40 chromatin markers in single cells. This study revealed increased epigenetic variation in immune cells with aging, providing unprecedented insights into the epigenetic basis of immune system changes over the lifespan.
Histone Modification Profiling
Building on the EpiTOF technique, Falquet et al. (2023) published a study in iSCience titled “Dynamic single-cell regulomes characterize human peripheral blood innate lymphoid cell subpopulations.” This study utilized single-cell RNA and ATAC sequencing to analyze the heterogeneity and regulatory factors of human peripheral blood innate lymphoid cells (ILCs). The researchers identified distinct transcriptional profiles and chromatin accessibility states within main ILC subsets, focusing on ILC2s and ILC progenitors (ILCPs). They discovered two distinct subpopulations each for ILC2s and ILCPs, with varying degrees of commitment and activation. Notably, ILC2a showed more pronounced ILC2 commitment and tissue-protective features, while ILC2b displayed a more migratory phenotype. For ILCPs, they identified a less mature ILCPa subpopulation and a more activated ILCPb subpopulation. This single-cell multi-omics resource from healthy donors demonstrates potential for inferring and predicting ILC alterations in disease conditions, possibly facilitating hypothesis-driven studies in patients without requiring extensive patient sample analysis.
DNA Methylation Analysis in Rare Cell Populations
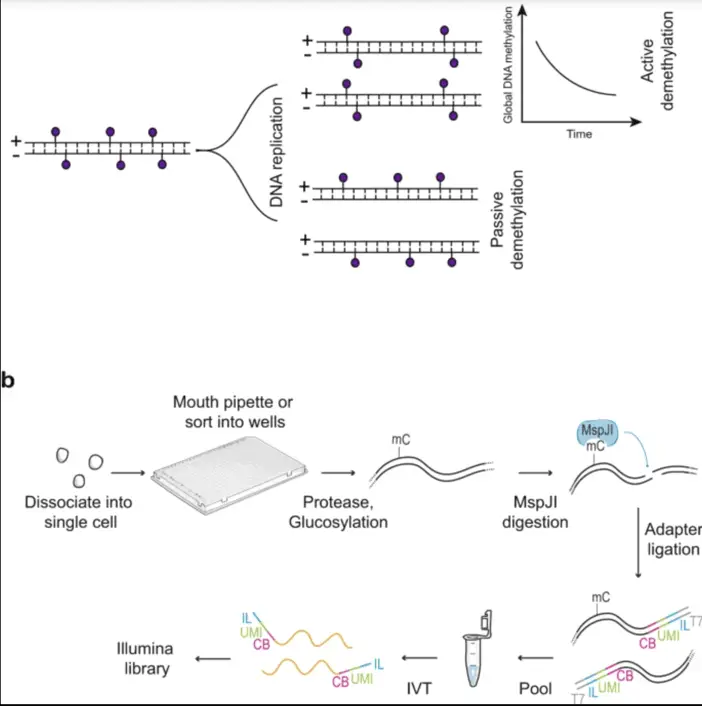
Mass cytometry has also been adapted for DNA methylation analysis at single-cell resolution. Sen et al. (2021) introduced a method called scMspJI-seq in Nature Methods, which combines restriction enzyme-based DNA methylation profiling with single-cell sequencing. While not directly using mass cytometry, this approach demonstrates the growing interest in single-cell epigenetic profiling.
Application in Tumor Immunology for Solid Cancer
In the context of tumor immunology, epigenetic profiling at single-cell resolution has provided valuable insights into the complexity of the tumor microenvironment. A study by Satpathy et al. (2019) published in Nature Biotechnology, titled “Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion,” used a combination of single-cell ATAC-seq and RNA-seq to profile epigenetic states of tumor-infiltrating immune cells in basal cell carcinoma.
While this study didn’t directly use mass cytometry, it demonstrates the power of single-cell epigenetic profiling in understanding tumor immunology. The authors identified distinct chromatin states associated with T cell exhaustion in the tumor microenvironment, providing potential targets for immunotherapy.
The integration of epigenetic profiling with mass cytometry and other single-cell technologies is rapidly advancing our understanding of cellular heterogeneity in complex tissues like tumors. As these techniques continue to evolve, they promise to provide even deeper insights into the epigenetic regulation of immune responses in cancer and other diseases, potentially guiding the development of new therapeutic strategies.
Epigenetics - where your DNA is just the script, but the real drama is in how it's read! I once heard an epigeneticist say, "DNA is like a piano. Epigenetics determines whether you play Chopin or chopsticks." It's fascinating to think that our experiences can 'tune' our genetic piano, influencing which genes are expressed. One study that tickled my funny bone showed that men who overate as children had sons who were less likely to die from diabetes. It's like your grandpa's big appetite was secretly preparing you for a healthier life! Who knew gluttony could be so... epigenetically beneficial? No need to check it out, here is everything to calm down your appetite : "We showed that individuals who were prenatally exposed to famine during the Dutch Hunger Winter in 1944–45 had, 6 decades later, less DNA methylation of the imprinted IGF2 gene compared with their unexposed, same-sex siblings." Yes, it is again a Dutch study....
Guillaume Beyrend